-
The Navigator helps you select and examine a single slice from a whole brain.
Brain image datasets are acquired using several imaging technologies, including magnetic resonance structural imaging and radionuclide functional imaging. In some cases datasets are gathered repeatedly over a time interval, in order to see how the brain changes, either by itself or in response to drug or surgical treatment.
- How is it possible to see the same part of the brain with different techniques or the same part across time? In each Whole Brain Atlas case, the datasets are carefully superposed, or shown "in register".
This permits precise comparison of data from the same slice plane taken from different image types or different time points.
Comparisons are even easier with the Java applet version of the Navigator, but your browser must support Java. We recommend Netscape 3.0 or later.
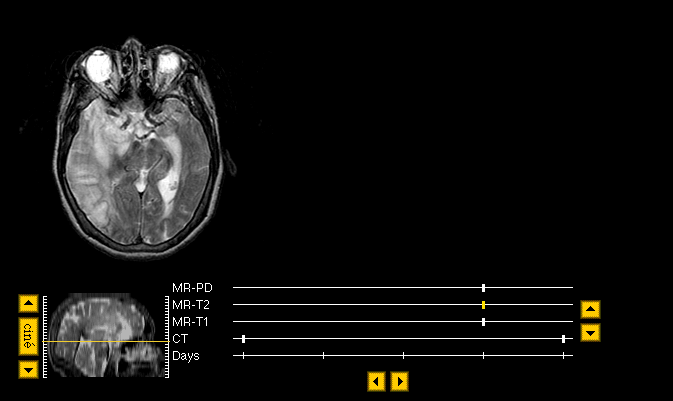
- Here is an example of the Navigator. The image at top is a T2-weighted MR in the transaxial plane. To learn more about what that means, check the Neuroimaging Primer, and the MR help pages. Click on the image to download a copy in the gif format. The subject's left is at right; this unfortunate convention has been widely adopted, and is used in the Atlas.
The Navigator's tools appear beneath the main image, and will be described in detail below.
- The location of the image in the whole brain dataset is shown as the yellow line in the sagittal image (the side view). The buttons to the left retrieve the next or previous image above or below the current image. The sagittal image is an "imagemap", so the slice can be directly chosen by clicking on the sagittal image at the desired slice location. The "cine" button retrieves a movie (MPEG).
As of August 20, 1997, to get an MPEG player, try ftp://ftp.netcom.com/pub/cf/cfogg/mpeg2. If you are running Windows95 (condolences), you might want to poke around http://www.windows95.com; last we checked, Win95 MPEG players can be found on this site at: http://www.windows95.com/apps/video.html.

- The list of image types available for the current case is shown at left. Abbreviations are expanded at the bottom of this help page. The time scale in this example is Days, at bottom left. The tickmark for the currently visible datset, MR-T2, is shown in yellow. The arrows at right change the image type; for example, clicking the up arrow will change the main image to a proton density image, MR-PD, and clicking the down arrow will jump to a T1-weighted MR image, MR-T1. The image dataset can also be directly selected by clicking on the tickmark. The arrows at bottom change the timepoint for, in this case, CT, which was acquired at two times separated by four days.
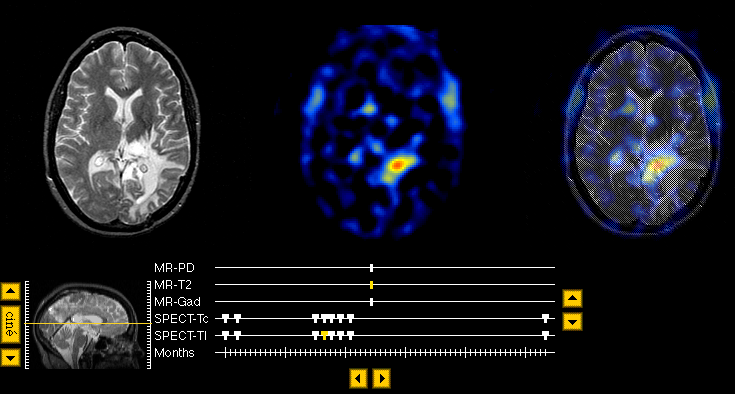
- Here is another example of a Navigator page, taken from one of the tumor cases . This Navigator page shows two images, T2-weighted MR, Thallium SPECT, and their overlay, indicated on the timelines by yellow tickmarks. These images are taken from a case in which 3 MR, 9 Thallium SPECT, and 9 perfusion SPECT datasets are available.
Note that some of the timeline tickmarks have a thick part and a thin part. Tickmarks with thick upper halves correspond to datasets sharing an overlay with the "current" dataset (the leftmost of the three slice images, represented on the timeline by the yellow, uniformly thin tickmark). Click on the thick part to see an overlay of the same "leftmost" image with the new dataset corresponding to that tickmark. The thin half retrieves the image type without an overlay. Again, the buttons at center, beneath the timeline, retrieve datasets from the next/previous time point. Each dataset is in registration with all others, so the same slice plane is sampled at each timepoint.